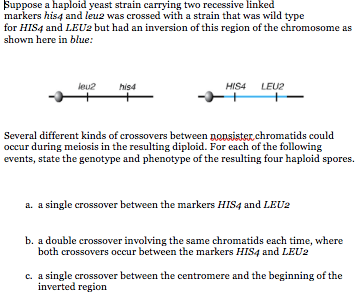
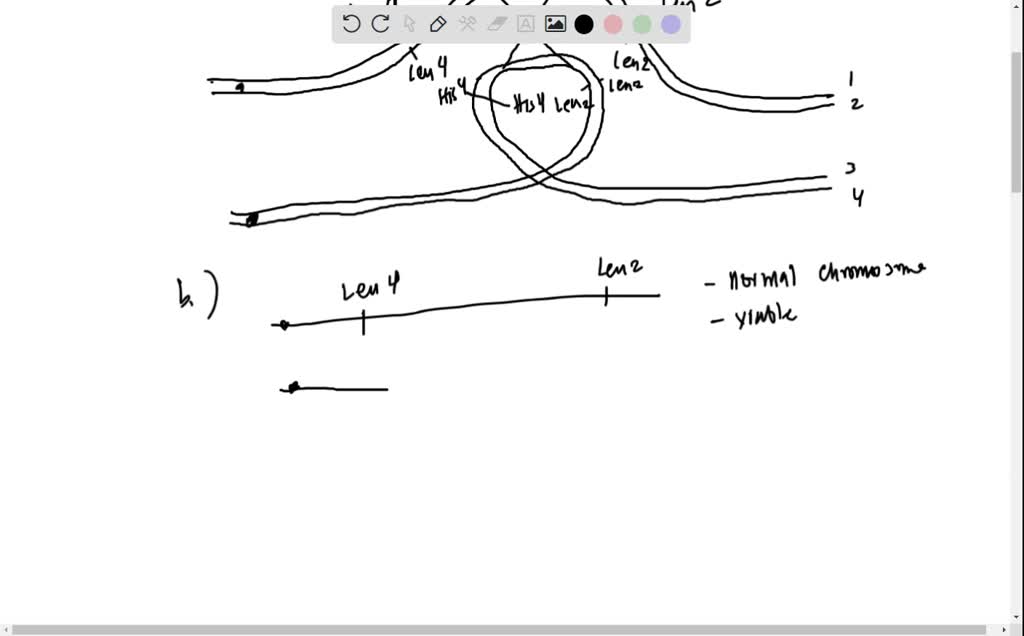
SOLVED: Map distances were determined for four different genes (MAT, HIS4, THR4, and LEU2) on chromosome III of the yeast Saccharomyces cerevisiae: HIS4-MAT 37 cM, THR4-LEU2 35 cM, LEU2-HIS4 23 cM, MAT-LEU2

Amazon.in: Buy Tout en un fran/maths/ang/his4 Book Online at Low Prices in India | Tout en un fran/maths/ang/his4 Reviews & Ratings
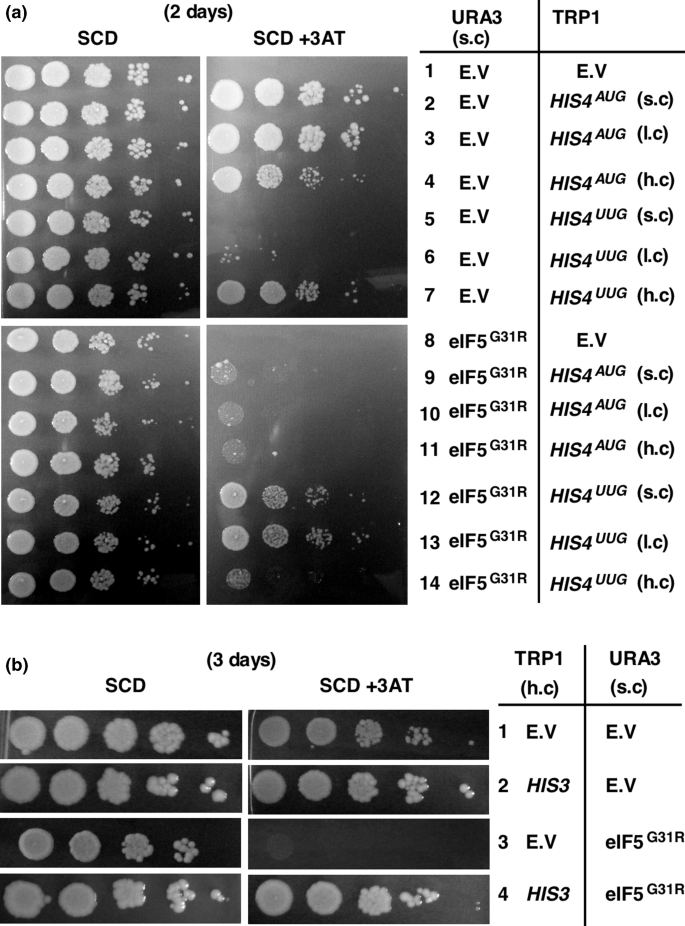
Fidelity of HIS4 start codon selection influences 3-amino-1,2,4-triazole sensitivity in GTPase activating protein function defective eIF5 | SpringerLink

Placement of the two adjacent genes, hIFNγ and HIS4, as part of the... | Download Scientific Diagram
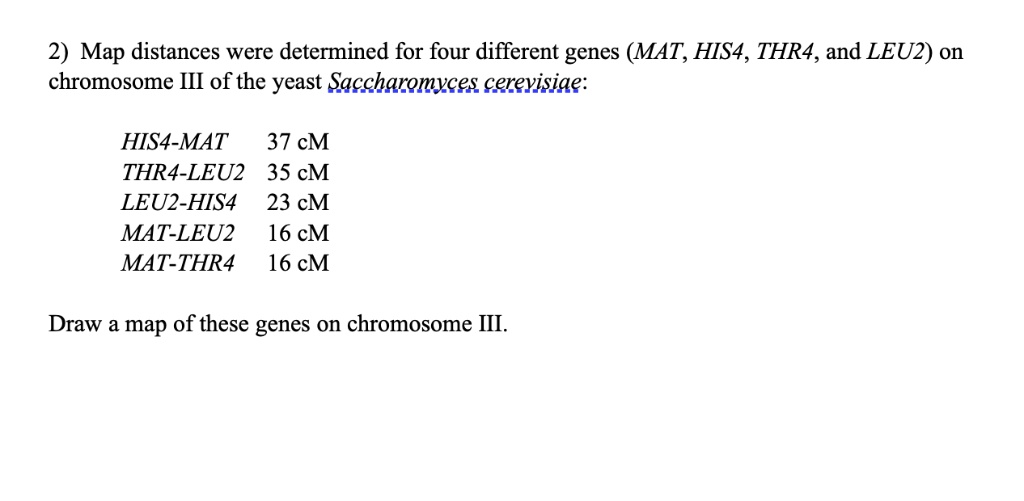

SOLVED: Map distances were determined for four different genes (MAT, HIS4, THR4, and LEU2) on chromosome III of the yeast Saccharomyces cerevisiae: HIS4-MAT 37 cM, THR4-LEU2 35 cM, LEU2-HIS4 23 cM, MAT-LEU2
![PDF] The conversion gradient at HIS4 of Saccharomyces cerevisiae. I. Heteroduplex rejection and restoration of Mendelian segregation. | Semantic Scholar PDF] The conversion gradient at HIS4 of Saccharomyces cerevisiae. I. Heteroduplex rejection and restoration of Mendelian segregation. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/b0a195533acf6089fe5ae45305300832ba362063/6-Table1-1.png)
PDF] The conversion gradient at HIS4 of Saccharomyces cerevisiae. I. Heteroduplex rejection and restoration of Mendelian segregation. | Semantic Scholar

Henchman HIS4, Midi Hi-Steps, Aluminium Garden Work Platform, Ideal to Cut 3.3m/11ft Hedges, 4 x Height Adjustable Legs : Amazon.co.uk: Garden

Meiotic recombination at HIS4 :: LEU2/his4-X :: LEU2-URA3 hot spot in... | Download Scientific Diagram

Recycling the HIS4 marker and eliminating the CRISPR/Cas9 system. a, b... | Download Scientific Diagram

![Somatostatin-14 [Des-Ala1, Des-Gly2, His4, 5, D-Trp8] peptide | Biorbyt Somatostatin-14 [Des-Ala1, Des-Gly2, His4, 5, D-Trp8] peptide | Biorbyt](https://www.biorbyt.com/pub/media/amasty/webp/catalog/product/cache/c687aa7517cf01e65c009f6943c2b1e9/s/o/somatostatin-14_[des-ala1,_des-gly2,_his4,_5,_d-trp8]_peptide_orb233911_1.webp)














